PharmCAT

Easy and popular: PharmCAT
PharmCAT is a bioinformatics tool used to analyze genetic variants from VCF files to identify pharmacogenomic haplotypes and predict how an individual may respond to certain medications.
In brief, it is used to:
-
Detect pharmacogenomic haplotypes (star alleles) in relevant genes.
-
Interpret variants based on guidelines from the Clinical Pharmacogenetics Implementation Consortium (CPIC).
-
Generate clinical reports describing potential drug response, efficacy, or risk of adverse reactions.
-
It is mainly used in clinical pharmacogenomics to support personalized medicine and drug therapy decisions based on a patient’s genetic profile.
Installation
Pharmacogenetics
sudo apt update
sudo apt install docker.io
docker --version
sudo systemctl start docker
Then you load it into docker
docker pull pgkb/pharmcat
Run it
Now that you have PharmCAT installed, you can analyze your patients' .vcf files to identify genetic variants that may affect their response to medications. This can be highly relevant in conditions such as hypercholesterolemia, where certain SNPs may impair the metabolism of statins and potentially lead to additional complications for the patient.
sudo docker run --rm -v $(pwd):/pharmcat/data pgkb/pharmcat pharmcat -vcf /pharmcat/data/pharmcat.example.vcf
If you have multiple files, you can process them all at once with:
for file in *.vcf
do
sudo docker run --rm -v $(pwd):/pharmcat/data pgkb/pharmcat \
pharmcat -vcf /pharmcat/data/$file
done
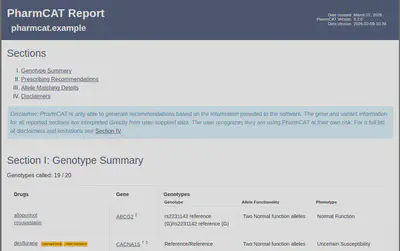
Finally, you will obtain a report like this, which allows you to evaluate the potential impact of specific variants on drug response. Keep in mind that each patient must be interpreted within their clinical context.